I love programming competitions and attempt to try every new site I find. Those sites will worth another post later, this current will be about a programming contest on CoderCharts I took part recently. It finished about two days ago and I didn’t do as well as I hoped, but did much better than I would have a year ago. Out of the 8 puzzles i have: 5 solved, 1 given up on for now, 1 haven’t even started and 1 attempted but run out of time . This last on is one Lemmings mating, one of the “hard” puzzles, and when I say run out of time I mean at 4am I decided that for the 4 ours then left to finish the competition I’d rather sleep than try to think of another optimization. Still, it bothered be and I did manage to write solution (with ~70% score, not too bad). Now I want to document the how did I get there so I won’t forget.
* Spolier warning. If you haven’t solved the problem and want to do it independently, do not read on. *
Path to a solution
Looking at the problem statement, there’s clearly a graph-related problem behind all the story of the poor creatures. When I’m not sure about not just what should be the solution, but what exactly is the right description the problem, I usually open The Algorithm Design Manual (ADM). It’s a great book, so far I couldn’t find a problem where it had nothing to say. It is a bit dated, however, and while sets me on the right path, there’s usually much more reading than that. In this situation it quickly revealed that the problem I face is finding the maximum clique in the graph describing the problem. Well, quickly learned that there’s a big can of worms waiting for me in there.
Maximum clique
So, to summarize, a clique is a group of vertices that are all connected to each other by edges. Got a maximal clique when it is not possible to add any other vertex in a way that everyone’s still connected. Finally the maximum clique is the largest maximal clique. Note how logical the notation is but still so easy to mix up. It is relatively easy to find maximal cliques (start from any vertex and add more connected vertices until there’s none left that is connected to all already selected). Finding the maximum clique (or any maximum clique since there could be more than one) is, however, a proper NP-complete problem.
The ADM only says something along the lines, that “well, then you just have to do a backtracking through all the graph”. Which is a great idea and simple to implement, I would had a result, but it is terribly inefficient because in a naive implementation one would check lots of cases which cannot possibly have the result. So fist thing to do is pruning, or eliminating paths that we are sure that it cannot improve what we already have. Wikipedia helps in finding just such a thing, called Bron–Kerbosch algorithm. Idea is quite simple (just wrap the head around recursive functions) and it can be implemented in Python following the pseudo-code version on Wikipedia. Here’s mine (not really optimized or anything).
def bronkerbosch(R, P, X, graph):
"""
Want all vertices from R, some from P and none from X
where the graph dict defines the connections
"""
if len(P) == 0 and len(X) == 0:
return R
candidates = P.copy()
max = set([])
for v in candidates:
res = bronker1(R.union([v]),
P.intersection(graph[v]),
X.intersection(graph[v]),
graph
)
if len(res) > len(max):
max = res
P.remove(v)
X.add(v)
return maxThis really worked and actually 3 out of 7 tests are passed, but the rest timed out. Got to find something better. Looking at more on Wikipedia, I was tempted to get on with the version of the algorithm that they said is “the fastest”, by J.M. Robson. It’s written up, but it is from the pre-MathJax era, so it’s just terrible work to figure out and keep up with all the strange notation. Also, it looks like a big collection of special cases and theoretical shortcuts. I’m sure it works, maybe I’ll come back to it later but for now I wanted to have a little bit more of an insight as well, so looked a bit more.
Vertex colouring
Since at this point I really caught up with the fact that maximum clique finding is an important problem, so went to Google Scholar to see what the academia has written. A lot, apparently. I was browsing though them in a kind of reverse-chronological order, so I could find the “newest” algorithms, since that should be the best. In the end a pattern emerged: most of the hard work can be done by employing a different technique, the vertex colouring. The two are connected because a clique in a graph G is an independent set in the complement graph of G (two vertices are connected in the complement of G if and only if are not connected in G), and vertex coloring is a good method of finding independent sets. What we want is labeling the vertices with the smallest number of different labels (“colours”) such that vertices with the same label are independent from each other. The number of different labels we have to use to do that helps pruning our clique finding algorithm by setting an upper bound how big clique it is possible to make from the vertices we have in a particular step.
As an example we can start out with a graph like this (taken from the original lemmings problem):
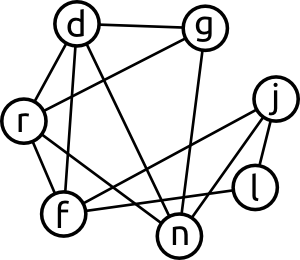
When running this graph through a vertex colouring algorithm, we would get something along this lines:
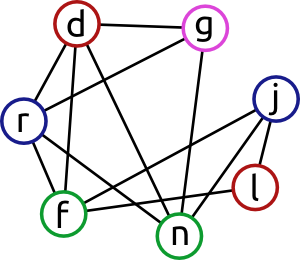
The blues are not connected to each other, neither are the greens and so on. In this case we need 4 colours to separate the graph into independent groups. Of course when programming one would use numbers instead of colours (thus often this called “numbering”): blue was actually “1”, red was “2”, … By adjusting the algorithm in terms of what is the order of the colours and the order of the vertices within each colour the previous algorithm can be improved by orders of magnitudes since we much better pruning. In the maximal clique every vertex would have different colour (though not all colours will necessarily be used).
Some notes on the algorithm to get started:
- Try to keep nodes in the order of neighbours in the current sub-graph. The smaller is good, because they can be quickly eliminated, reducing the test-space more.
- Go from colors with fewer members to more members, similar reasons.
- Can use the color number + the current clique size for better pruning: if my largest clique so far has Q elements, my current clique has P and the tested vertex comes from the highest colour number N, then in case of N + P <= Q it is futile to go on, I cannot improve the result and time to backtrack. Less efficient methods use the C number of elements in the current candidate set instead of N, and since N<=C, the pruning bound set by C+P is more “loose” letting more tests to be done than necessary.
For example, the final solution of this given problem is the set of vertices highlighted by the edges is shown on the next picture, and it is done in only 4 steps or so…
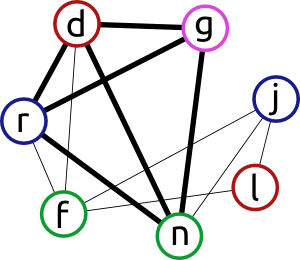
Different methods
In the papers I found a few different colouring schemes, and most of the improvements seem to be made in:
- initial ordering, or
- ordering within each of the colours, or
- better upper-bound estimation for the possible clique size from the available colour information.
The hardest things about reading these papers is that often the pseudo-code is a big mess, there are plenty of typos and the examples are quite scarce.
In the end I used algorithms mixed from two different papers:
Tomita, Akutsu and Matsunaga, Efficient Algorithms for Finding Maximum and Maximal Cliques: Effective Tools for Bioinformatics, 2011 (link) This has a few different algorithms and the most effective one looked too complicated and I went with MCQ, one of the improved but not perfect on. Now 6/7 tests pass, that’s got to be the right path. According to benchmarks the coloring takes up the most time
Segundo, Rodriguez-Losada and Jimeneza, An exact bit-parallel algorithm for the maximum clique problem, Computers & Operations Research, Volume 38, Issue 2, 2011, pp.571-581 (doi:10.1016/j.cor.2010.07.019). Their algorithm is a variation on the previous one and I’m not using it to it’s full potential (wonder how fast a C implementation would be). Also, I wonder if I misunderstood something but the original form of the algorithm failed on me. There are nodes that don’t need to have colour information because it does not matter at that stage of the algorithm. Their version of colouring then seems to remove those vertices from the colouring function output. This does not matter for their example (it is correct) but in my tests it gives wrong result (the algorithm finishes too early). So, mixed in from the previous paper that I keep all nodes whether they are coloured now or not, because later they might be, it works like a charm. And about a factor of 2-3 faster than before, enough to pass all 7 tests on CoderCharts and with a reasonable score. I think the real improvement is not really in the pruning (this version seems to check a few more nodes than the previous) but somehow the colouring function is faster – that might tell something about my Python skills too, though.
Lessons
This could probably be solved with other algorithms as well, I would be curious to see others’ code / hear their stories. What I took home from this:
- Know your problem and know where to look for solutions. The best is to study and practice a lot.
- Often pre-processing the data is a crucial step to the solution. Since there’s usually a memory/speed tradeoff, it is worth experimenting: if and algorithm is too slow, what could be prepared and stores to exchange calculating something with a mere data lookup?
- Different algorithms have bottlenecks at different sections. E.g. in this case there is the data preparation, the number of steps to test possible solutions and the time it takes for re-colouring. On the path to the final solution I had algorithms inefficient in each of the sections.
- Test cases are important. The example in the problem setting and the examples in all the papers are too small to be helpful for optimization. The “bonus” test case on problem site, however , is just too big. Actually, even my passed algorithm is too slow to have a result for it in reasonable time. So, if there’s no provided test case, make your own
- The example cases I generated are much smaller than the 7 tests I need to pass, but they are still slower. Every paper said that maximum clique finding has very different running time for different (random) graphs. I presume that some of the test graphs are in fact specially prepared. Also, some of the changes in code that made my own test cases faster (sometimes 20%) often failed more tests on the web. The message is that coding competitions like this are set towards finding the optimal solution for their own tests – whatever your winning solution might be, it is not necessarily the best overall solution and if I ever come across a similar problem in real life setting I likely to have to do differently.
- Most of the Python optimization advice I found is outdated and sometimes outright hurting the performance. Do plenty of performance testing. Start optimizing the big slow downs, the small shortcuts rarely worth the time in competition setting.
- Language choice matters but does not matter that much. I’d think Python actually pushes me to be more efficient because many things are slower. The code on the other hand is more readable in the end (ideally).
- Someone has to update Wikipedia. :)
I will share the code later, just need to wait until some time has passed after the competition.